Fibroblast Activation Validated Solution
3D Fibroblast Activation Workflow with RASTRUM
Start from a validated workflow designed to model fibroblast activation and stromal signaling in a controlled context of use.
The Fibroblast Activation Validated Solution supports researchers studying fibroblast activation, extracellular matrix remodeling, and cellular crosstalk in biologically relevant 3D environments.
Model overview
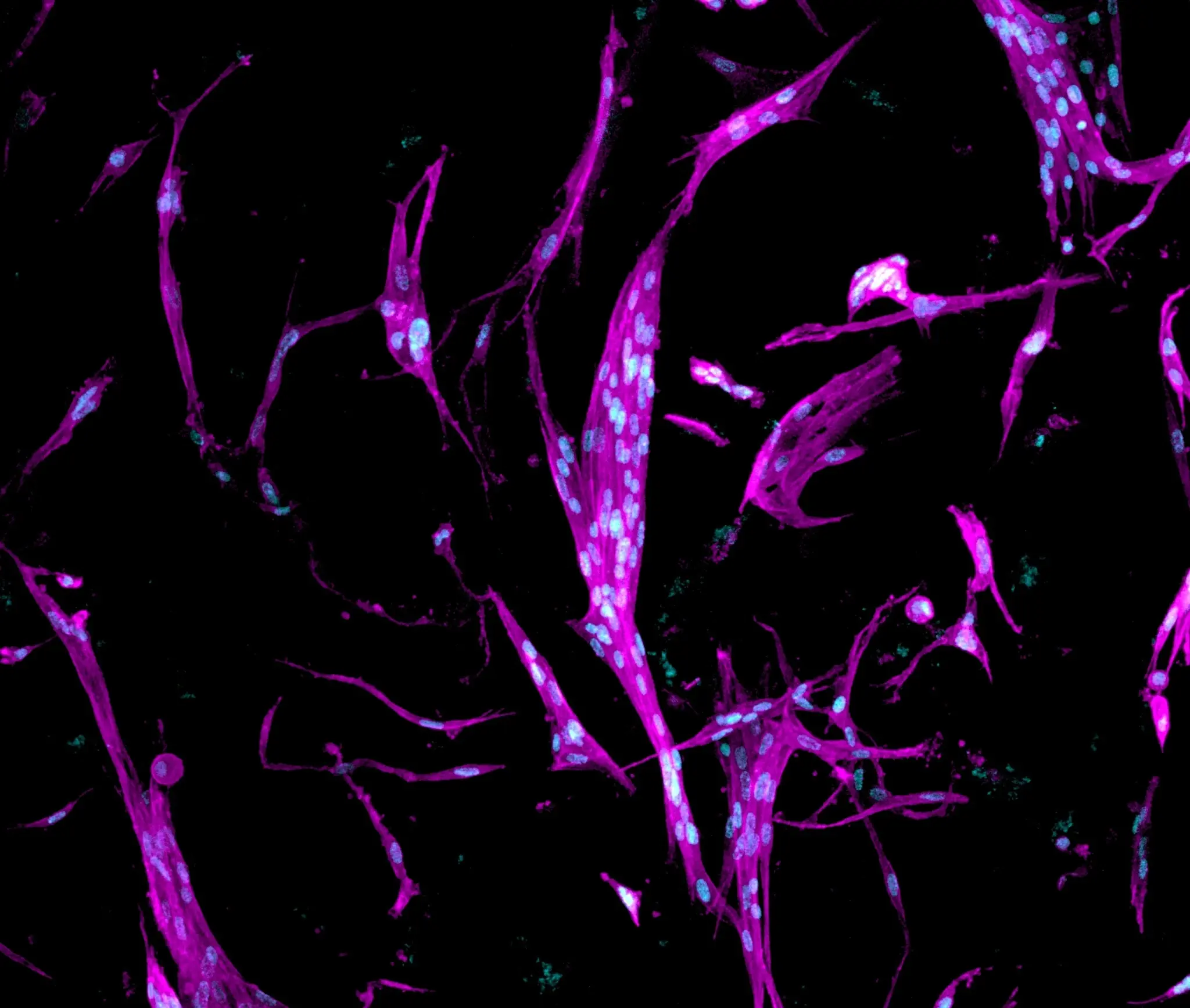
RASTRUM matrices maintain fibroblasts in an activatable state
Fibroblasts play essential roles within the tissue microenvironment through extracellular matrix remodeling and signaling. In solid cancers, fibroblast activation can modulate tumor cell biology and inhibit the efficacy of drug treatments, including immunotherapies. Traditional 2D culture does not recapitulate the in vivo extracellular matrix and can drive uncontrolled activation, whereas controlled 3D environments can maintain fibroblasts in a non-activated state until deliberately stimulated.
Context of use
This validated workflow is designed for researchers who need a controlled 3D model of fibroblast activation for mechanistic studies, signaling analysis, and translational microenvironment research. It supports fibroblast monoculture modeling, induction of activation through TGF-β stimulation, and development of fibroblast-containing co-cultures for studying cellular crosstalk.
-
Create 3D monoculture fibroblast models with in vivo-like network morphologies that maintain fibroblasts in a non-activated state
-
Study fibroblast activation via addition of TGF-β and assess strategies to perturb this activation status
-
Develop 3D co-cultures of cancer cells and fibroblasts to model tumor microenvironment-like biology
What this model demonstrates
This Validated Solution is designed to support interetable, biologically relevant fibroblast activation studies in a controlled 3D context
-
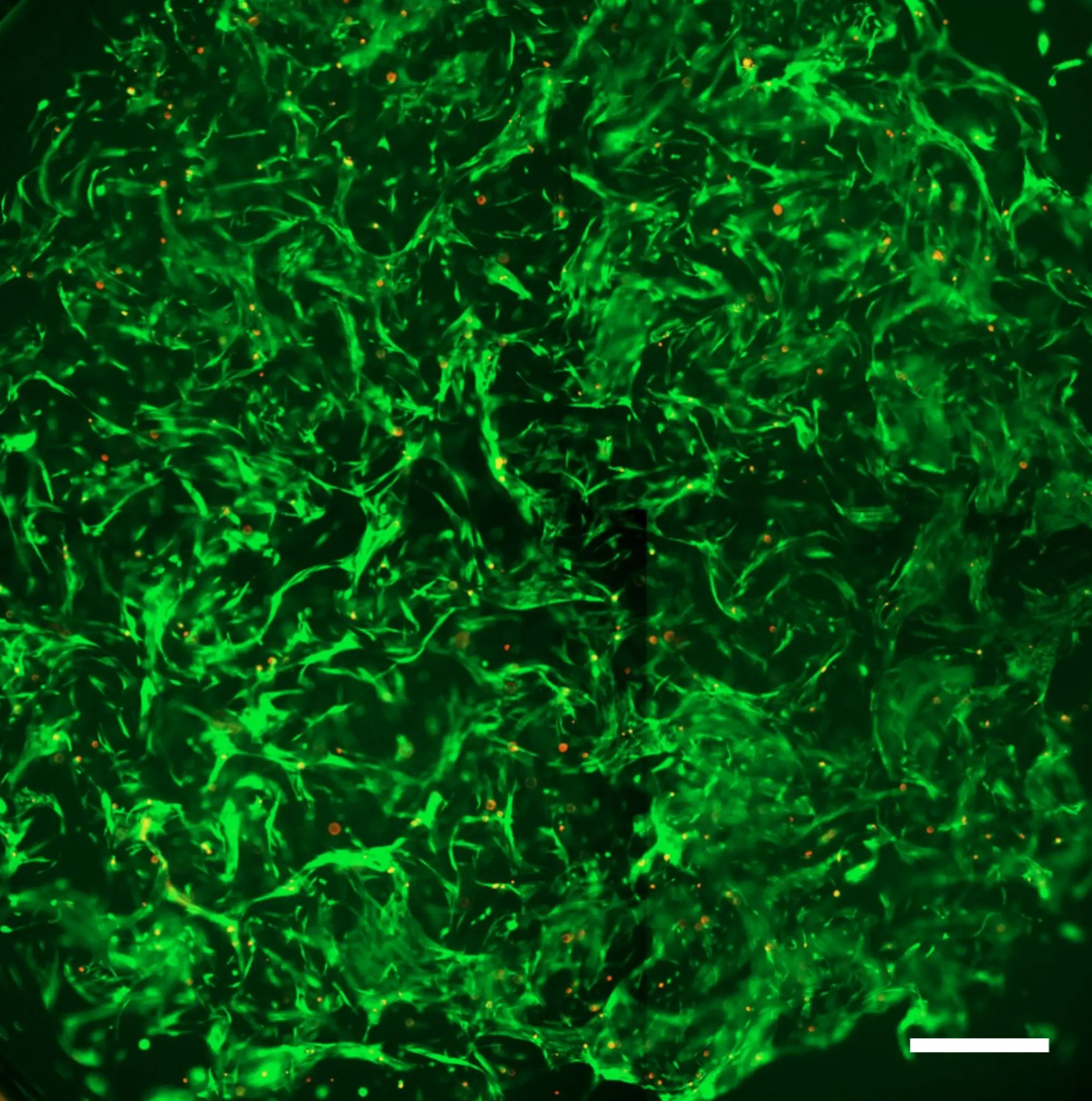
Fibroblasts can be maintained in 3D while preserving an activatable state
The workflow supports in vivo-like fibroblast morphologies without uncontrolled baseline activation. -
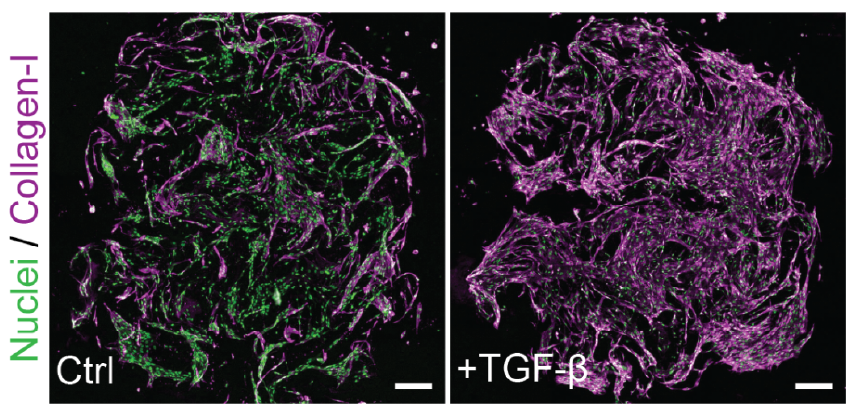
TGF-β drives a measurable activation response
Collagen I staining and image-based quantification, together with COL1A1 expression, confirm induced fibroblast activation. -
The model supports broader stromal and co-culture applications
The workflow provides a starting point for studying fibroblast signaling, extracellular matrix remodeling, and more complex fibroblast-containing co-culture systems relevant to tumor microenvironment biology.
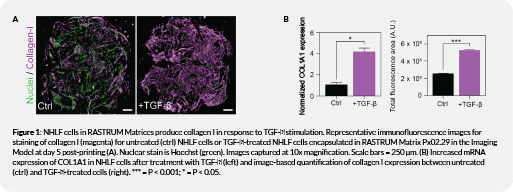
Molecular characterization
TGF-β induces measurable fibroblast activation in RASTRUM matrices
In this 3D fibroblast activation model, NHLF cells treated with TGF-β showed a marked increase in collagen I relative to untreated cells. Gene expression analysis further confirmed fibroblast activation, with treated cells showing an approximately fourfold increase in COL1A1 mRNA expression. Collectively, these data show that TGF-β treatment induces fibroblast activation in the RASTRUM model.
Key workflow specifications
Defined model parameters support reproducible execution across runs and downstream analyses.
- Model format: Imaging Model
- Matrix: Px02.29 (1.1 kPa, RGD, YIGSR, GFOGER, HA)
- Cell density: NHLF: 10M/mL
- Drug treatment: TGF-B day 4 post-print, 24 hr exposure
Get the detailed workflow package for this Validated Solution, including build parameters, reagent and materials requirements, operational guidance, and the supporting validation framework used for this defined context of use.
- Complete Protocol Pack
- Reagents and materials list
- Experimental workflow guidance
 Browse Validated Solutions
Predefined blueprints for reproducible 3D data in a defined context of use
Browse Validated Solutions
Predefined blueprints for reproducible 3D data in a defined context of use
 Explore Discovery Mode
Define and refine 3D cell models to match your biology and readouts
Explore Discovery Mode
Define and refine 3D cell models to match your biology and readouts
 Connect with Discovery Services
Expert support to optimize, validate, and integrate 3D models into discovery workflows
Connect with Discovery Services
Expert support to optimize, validate, and integrate 3D models into discovery workflows