PDAC Validated Solution
3D PDAC Tumor-CAF Co-culture Workflow with RASTRUM
Start from a validated pancreatic cancer co-culture workflow optimized for reproducible drug responses and tumor-stromal interaction studies.
The PDAC Validated Solution provides a repeatable 3D model with defined build parameters, biological and functional validation, and workflow guidance aligned to translational research applications.
Model overview
The RASTRUM PDAC Validated Solution recreates tumor-stroma interactions by combining pancreatic cancer cells with cancer-associated fibroblasts in a defined 3D matrix environment. To capture broad coverage of key PDAC signaling pathways, the workflow includes either PANC-1 or AsPC-1 cancer cells paired with primary PDAC CAFs , enabling comparison of drug efficacy across primary and metastatic contexts.
Context of use
This validated workflow is designed for researchers who need a reproducible 3D PDAC tumor-stroma model for therapeutic response studies in a defined context of use.
- Evaluate therapeutic efficacy in the presence of tumor-stroma crosstalk
- Investigate stromal contributions to tumor progression and drug resistance
- Identify targetable resistance mechanisms in the tumor microenvironment
- Screen pathway-targeted therapies and combination treatments
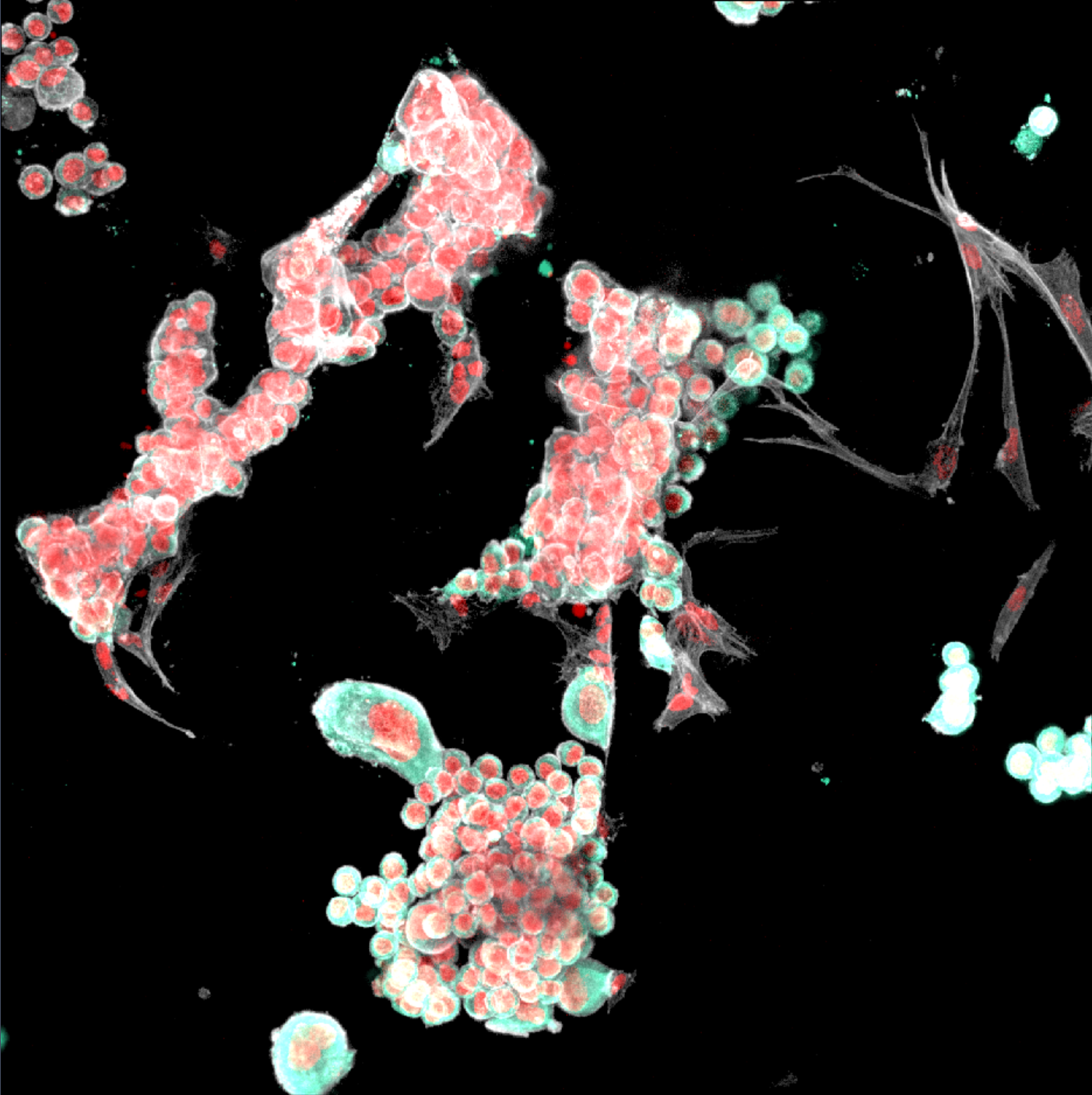
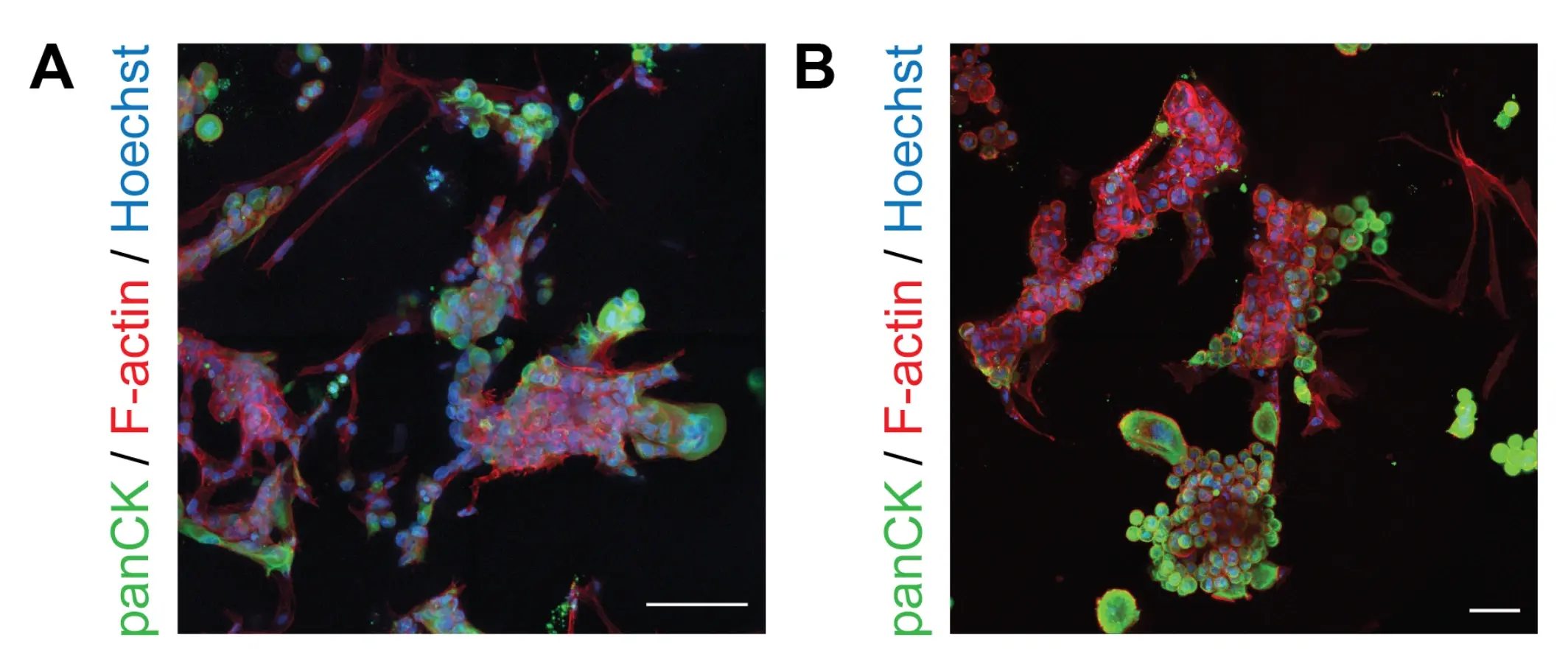
Image depicts immunofluorescence assessment of cancer cell–CAF interactions in RASTRUM optimized models. Positive immunofluorescence staining for pan cytokeratin (cancer cell marker, green) alongside phalloidin staining for actin filaments (red) after 7 days in culture. (A) PANC-1/CAF and (B) AsPC-1/CAF co-cultures.
What this model demonstrates
The PDAC Validated Solution is designed to generate interpretable, decision-grade insights in defined PDAC tumor-stroma contexts.
-
Stromal signaling is context-dependent
Distinct pathway enrichment profiles between PANC-1 and AsPC-1 highlight how tumor genotype shapes tumor-stroma crosstalk and downstream resistance mechanisms. -
CAF co-culture drives chemoprotection
AsPC-1 cells co-cultured with CAFs demonstrate reduced chemotherapy sensitivity, consistent with stromal-mediated protection. -
The model supports combination therapy screening
The workflow can be used to evaluate therapeutic strategies that counteract CAF-mediated drug resistance or remain effective in the presence of tumor-stroma signaling.
Molecular characterization
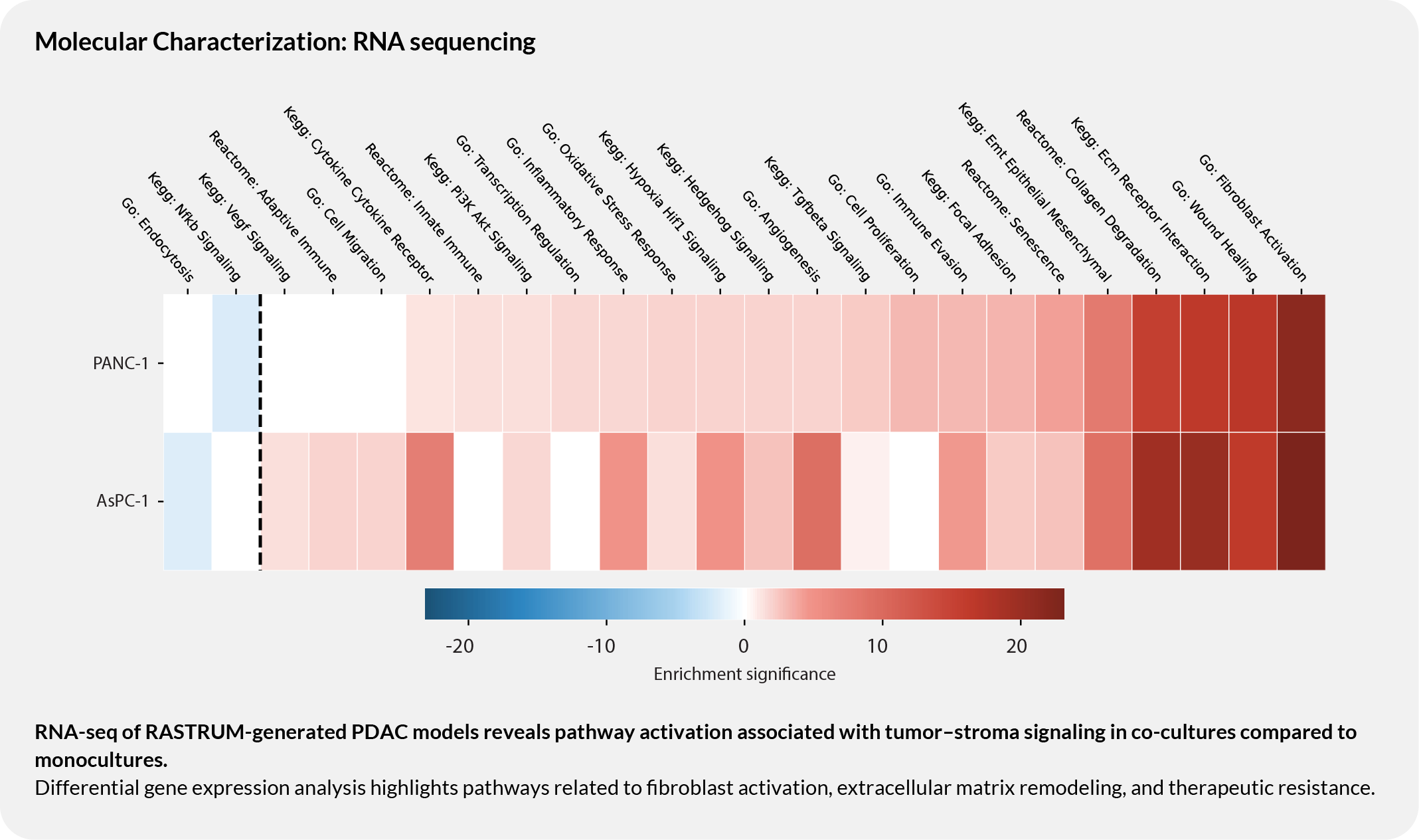
RNA-seq reveals pathway activation associated with tumor-stroma signaling
Differential gene expression analysis highlights pathway enrichment related to fibroblast activation, pro-tumourigenic processes, and therapeutic resistance. Distinct pathway profiles between PANC-1 and AsPC-1 co-cultures support the use of this workflow in biologically rich PDAC contexts.
Functional validation
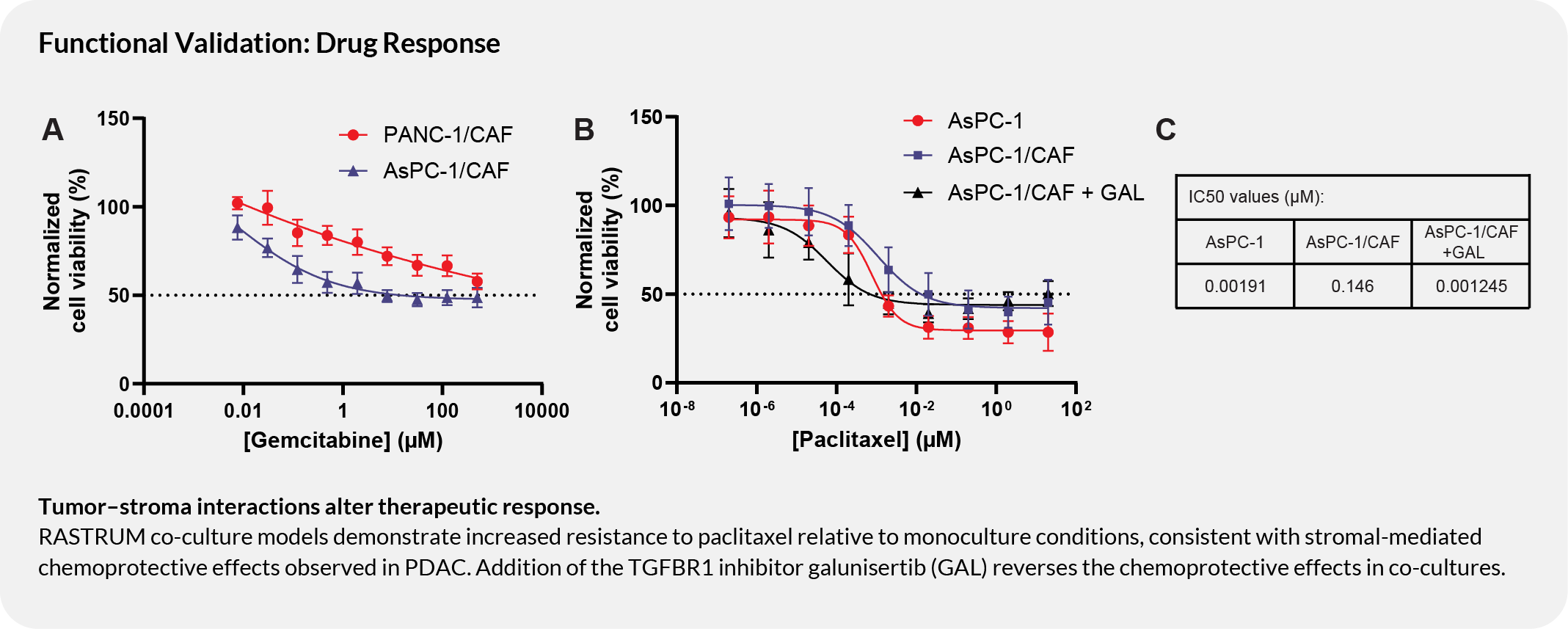
Tumor-stroma interactions alter therapeutic response
Co-culture models demonstrate increased resistance to paclitaxel relative to monoculture conditions, consistent with stromal-mediated chemoprotective effects observed in PDAC. Addition of the TGFBR1 inhibitor galunisertib reverses the chemoprotective effect in co-cultures, supporting use of this model for combination therapy evaluation.
Key workflow specifications
Defined model parameters support reproducible execution across runs and downstream analyses.
-
Model format: Imaging Model
-
Matrix: Px03.84 (~3 kPa, RGD, YIGSR, GFOGER, HA, full-length fibronectin)
-
Cell density: Cancer cells 2M/mL; CAFs 10M/mL
-
Co-culture ratio: 1:5 tumor:CAF
-
Drug treatment: Day 4 post-print, 72 hr exposure
-
Reference compounds: Gemcitabine, paclitaxel, galunisertib
-
Downstream analyses: Viability assessment, live/dead and brightfield imaging, drug screening, optional RNA preparation for sequencing

Get the detailed workflow package for this Validated Solution, including build parameters, reagent and materials requirements, operational guidance, and the supporting validation framework used for this defined context of use.
- Complete Protocol Pack
- Reagent and materials list
- Experimental workflow guidance
 Browse Validated Solutions
Predefined blueprints for reproducible 3D data in a defined context of use
Browse Validated Solutions
Predefined blueprints for reproducible 3D data in a defined context of use
 Explore Discovery Mode
Define and refine 3D cell models to match your biology and readouts
Explore Discovery Mode
Define and refine 3D cell models to match your biology and readouts
 Connect with Discovery Services
Expert support to optimize, validate, and integrate 3D models into discovery workflows
Connect with Discovery Services
Expert support to optimize, validate, and integrate 3D models into discovery workflows